Nanoimagerie : dynamique cellulaire et biologie quantitative
Our web site has moved to this address: https://lob.ip-paris.fr
Permanent staff
Antigoni Alexandrou (senior research scientist - CNRS)
Cédric Bouzigues (assistant professor - Polytechnique)
Nicolas Olivier (research scientist - CNRS)
Rivo Ramodiharilafy (assistant engineer - Polytechnique)
Position proposals |
|
Ph. D. Thesis/Internship proposal: In vivo detection of reactive oxygen species using multifunctional luminescent nanoparticles PhD Thesis/Internship proposal: Monitoring complex kidney pathophysiological processes in quantitative optical imaging in glomeruli-on-chip devices
Applications from highly talented and motivated candidates are encouraged. |
Research topics
Super-resolution Microscopy by Localization of Single Molecules
We work on nanoscale imaging using localization microscopy: This method relies on continuous imaging of blinking individual fluorescent molecules to ensure that only a subset of molecules per frame are visible and can be localized accurately. Adding up the localizations allows the reconstruction of a high resolution image. A number of methods ghave been developed such as PALM, STORM, or PAINT, which differ in the way the blinking is achieved. At LOB, we focus on developing STORM-like methods, where the fluorophores used are stanard organic dyes (such as Cyanine dyes, FITC, Rhodamine,...) and blinking is achieved by fine-tuning the chemical environment. In particular, our work on method development focuses on a few different topics:
- Improving the resolution of single colour towards the molecular scale
- Developping robust protocols for multicolor imaging
- Developing new methods for extracting reliable measurements from raw data
We also want to apply this powerful imaging method to answer biological questions. In particular we study the nano-organization of centrioles: They are evolutionarily conserved barrel-shaped organelles about ~ 500 nm in height and with an outer diameter of ~250nm. Centrioles are composed of more than 150 different proteins, and are essential for the organization of pericentriolar material and the formation of functional centrosomes, which in turn serve as the main microtubule organizing centre in most cycling cells and also as the precursor to cilia formation in quiescent cells or flagella. Because of their fundamental role in cell division to ensure proper separation of genetic material, centrosomes and centrioles play an important role in health and disease but so far the direct mechanisms governing centriole biogenesis and organization remain unclear. In particular although most of the constituting proteins have been identified using knock-outs, how they are scaffolded and in which order remains unknown.
Nanoparticles
We develop non-blinking single-molecule labels and sensors based on lanthanide-ion doped nanoparticles and use them for studying toxin-cell interaction and toxin receptor confinement in the cell membrane, and hydrogen peroxide production in signaling processes, respectively. We also develop and use microfluidic devices to produce asymmetric stimulation of cells and induce controlled deoxygenation of red blood cells in microfluidic droplets.
These nanoparticles are particularly attractive for single-molecule tracking applications (Nano Lett. 2004, Phys. Rev. Lett. 2009): they are synthesized directly in water, present high photostability and no blinking, narrow emission linewidths independent of nanoparticle size, and long excited state lifIntracellular H2O2 detection using Eu-doped nanoparticlesetimes which are useful for retarded detection schemes and FRET applications (J. Phys. Chem. B 2006). We furthermore discovered that Eu-doped nanoparticles are oxydant probes and can be used for quantitative time-resolved intracellular H2O2 detection with spatial resolution (Nat. Nanotech. 2009, Chem. Biol. 2014).
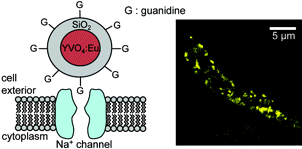
We demonstrated that these nanoparticles are detectable individually and that their size can be accurately determined from their luminosity (Appl. Phys. Lett. 2006). We have shown that nanoparticles functionalized with guanidine groups specifically target sodium channels and can be imaged individually on living cardiomyocytes (Nano Lett. 2004). We then implemented a more general nanoparticle-protein coupling scheme and counted the number of proteins coupled to individual nanoparticles (J. Am. Chem. Soc. 2007).
Nanoparticles labeling sodium channels on the membrane of a frog cardiomyocyte
Single receptor tracking in living cells to probe membrane nano-organization
We now use peptidic toxins (family of oligomerizing and pore forming toxins) labeled by Eu-doped nanoparticles to investigate their interaction with cells: binding to receptors on the cell membrane and receptor motion inside membrane microdomains. Novel single-molecule trajectory analysis techniques based on Bayesian inferences can extract the force field felt by the receptor (Phys. Rev. Lett. 2009, Biophys. J. 2010, 2012). These toxins can thus also be viewed as tools for investigating the membrane organization.
Quantititative Reactive Oxygen Species (ROS) signaling monitoring through nanoparticle imaging
Low concentrations of reactive oxygen species and, in particular hydrogen peroxide (H2O2), mediate numerous signaling processes in the cell. We measured the differences in the timing of intracellular H2O2 production triggered by different signals using Eu-doped nanoprobes (Nat. Nanotech 2009). We are now using this novel sensor to investigate the role of receptor transactivation and asymmetric stimulation in migration and contraction processes (Chem. Biol. 2014). 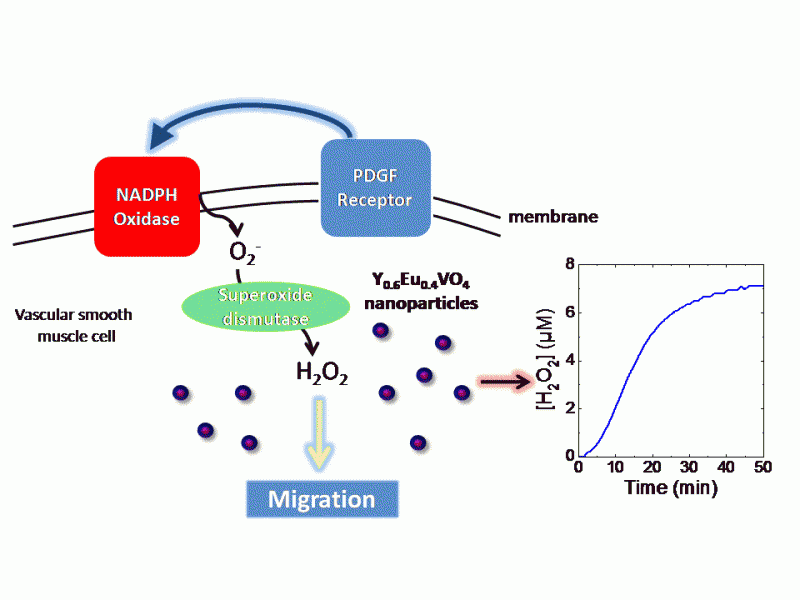
Dynamics of sickle red blood cells in microfluidic channels
Microfluidics offers the opportunity of generating controlled model environments reproducing physiological conditions while using minimal amounts of sample solutions. In particular, microfluidic channels are ideal for the study of blood cells in conditions approaching those of the blood flow.
We investigate the dynamics of red blood cells in the case of sickle cell disease, a genetic disease involving a single point mutation of the oxygen-carrying protein, hemoglobin. This mutation induces hemoglobin polymerization upon de-oxygenation which leads to a decrease in the deformability of red blood cells. This in turn leads to painful, vaso-occlusion episodes, organ damage and a shortened lifespan. We have implemented an innovating microfluidic platform that can generate cycles of oxygenation-deoxygenation close to the physiological ones to study this disease at the single cell level. Using polarization microscopy, we detect the hemoglobin polymerization by the birefringence it induces and study the behavior of numerous single red blood cells and the effect of important parameters and inhibitor molecules. This study should bring insight into the molecular mechanisms of the disease and help develop new therapeutic approaches.
Application to biomolecule ultra-sensitive detection in vitro
Lanthanide-based nanoparticles can be used as labels for immunassays (ELISA, Lateral Flow Assays,...) to significantly lower their detection thresholds. These applications have been pioneered in the team and are currently developed in collaboration with the spinoff/startup LumediX.
Former members :
Mouna Abdesselem (Ph.D. student 2012-2015)
Rachid Rezgui (Ph. D. student, 2010-2013)
Paul Abbyad (Post-doc, 2008-2012)
Markus Schöffel (Ph. D. student, 2009-2012)
Silvan Türkcan (Ph. D. student, 2007-2010 and post-doc 2011-2012)
Thanh-Liêm Nguyên (Ph.D. student, 2006-2009)
Didier Casanova (Ph. D. student, 2004-2007)
Egidius Auksorius (Marie-Curie exchange Ph. D. student, 2004)
Undergraduate students : G. Demangeon (2006), T. Amirtha (2005), T. Kourkoutsaki
Collaborations :
T. Gacoin, J.-P. Boilot, Lab. of condensed matter physics (Ecole Polytechnique, Palaiseau)
J.-M. Allain, Lab. de Mécanique des Solides (Ecole Polytechnique, Palaiseau)
M. Popoff, Unité Bactéries Anaérobies et Toxines (Institut Pasteur, Paris)
J.-B. Masson, M. Vergassola, Physics of Biological systems (Institut Pasteur, Paris)
P.-L. Tharaux, INSERM U970, Centre de Recherche Cardiovasculaire de Paris-PARCC