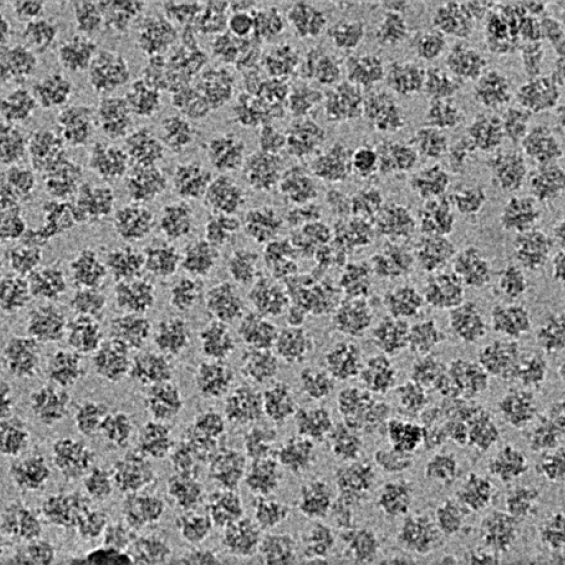
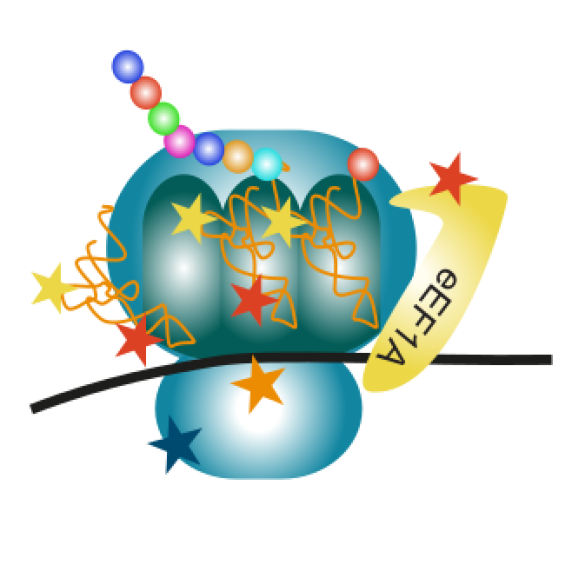
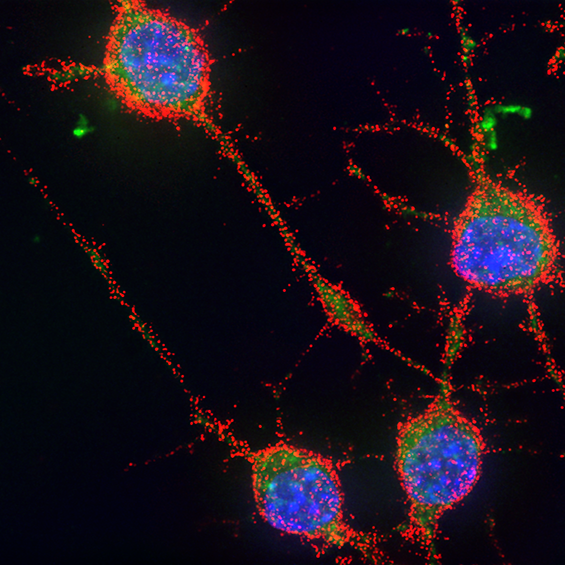
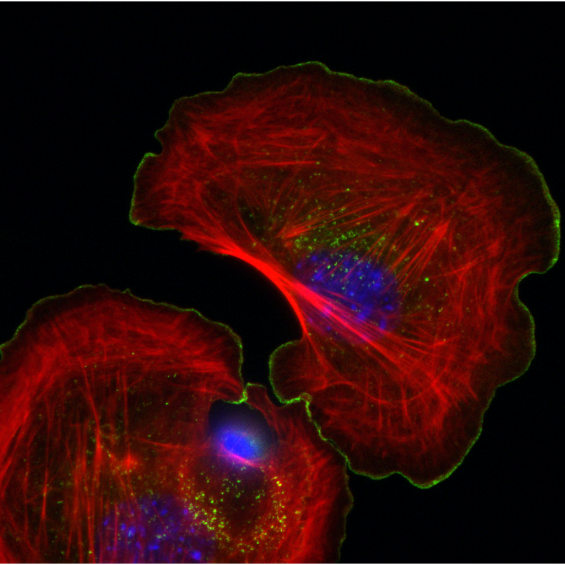
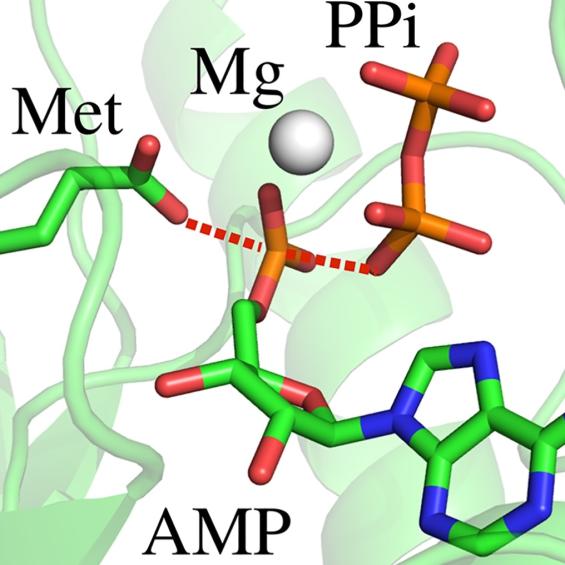
BIOC décrypte les processus cellulaires et moléculaires fondamentaux du vivant. En intégrant les niveaux atomiques, supra-moléculaires et cellulaires, BIOC explore des domaines tels que la synthèse protéique ribosomale, la migration et la signalisation cellulaires pour éclairer la compréhension de pathologies complexes, notamment le cancer et les maladies neurodégénératives, et pour mieux comprendre l’évolution.